Early detection of concerning SARS-CoV-2 variants through data analysis
Understanding the Evolution of SARS-CoV-2 Variants
Since the emergence of the SARS-CoV-2 pandemic, several variants of the virus have been identified and classified by the World Health Organization (WHO) as Variants of Concern (VOCs). These variants are considered significant because they may exhibit changes that increase their transmissibility, lead to more severe disease, reduce the effectiveness of vaccines, or place additional strain on healthcare systems. The ability to detect these variants early is crucial for public health planning and response.
The Role of CoVerage in Genomic Surveillance
A groundbreaking web platform called CoVerage has been developed to support genomic surveillance of the SARS-CoV-2 virus. This tool enables rapid computational identification and characterization of potential Variants of Interest (pVOIs), often with a lead time of nearly three months before they are officially designated as VOCs or related categories. By predicting the virus's ability to escape existing immunity from prior infections or vaccinations, CoVerage offers valuable insights into future outbreaks.
Researchers led by Alice McHardy have demonstrated the effectiveness of this system through a comprehensive analysis published in Nature Communications. Their work highlights the importance of early detection in vaccine development, ensuring that vaccines remain effective against emerging variants.
Key Components of the CoVerage System
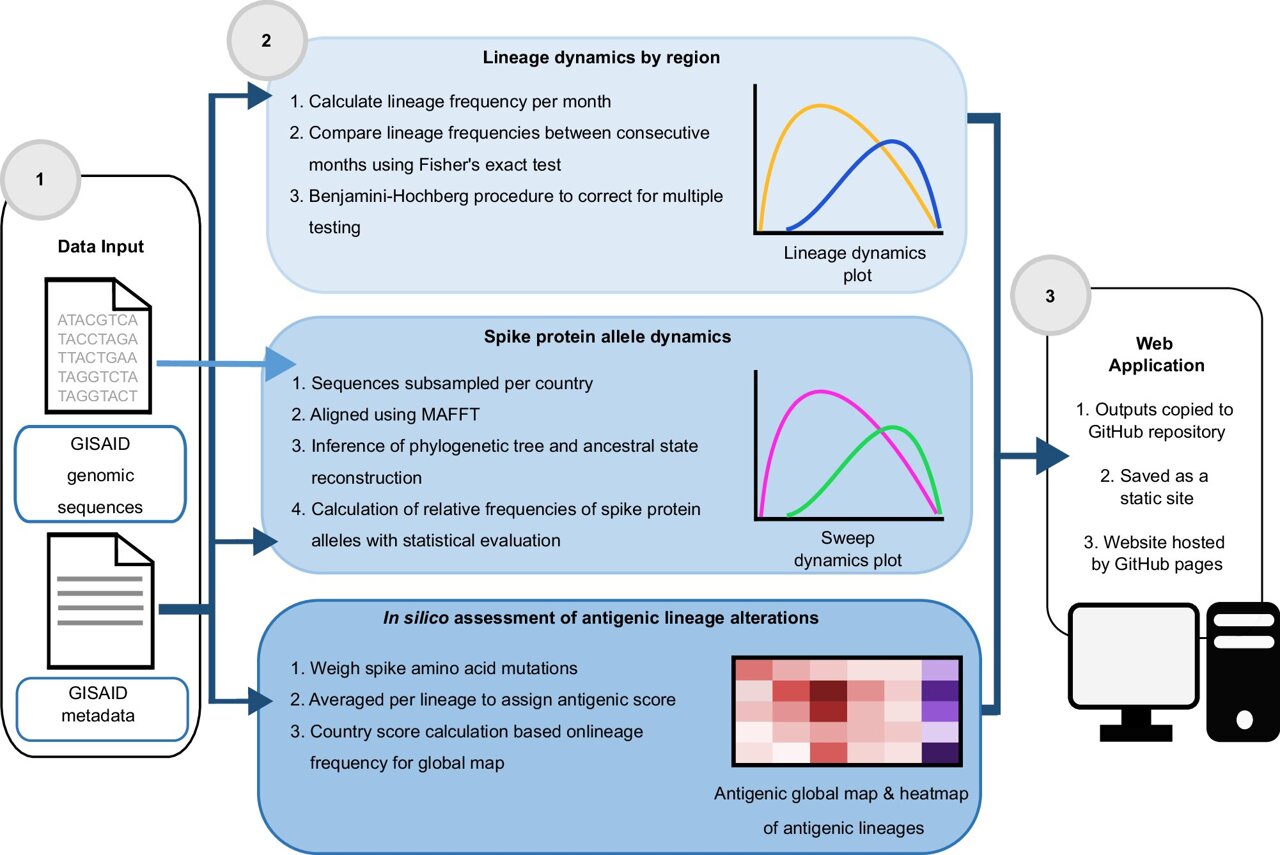
The CoVerage system utilizes a matrix based on observations from long-term developments of certain influenza viruses, such as influenza A H3N2. This matrix connects important genetic changes in the virus to its functional properties. The researchers focus particularly on changes in the spike protein, which is critical for the virus's ability to attach to human cells and is a primary target for vaccines and therapies.
Data for CoVerage is sourced from the GISAID virus genome database, a global initiative that facilitates the rapid exchange of data on priority pathogens. As of March 2024, GISAID had over 16.5 million SARS-CoV-2 sequences available, providing a rich dataset for analysis.
Analyzing Viral Strains and Antigenic Changes
CoVerage analyzes SARS-CoV-2 genome data by country of origin, tracking strain dynamics and antigenic changes. A statistical method is employed to identify viral strains that have significantly altered their immune escape capacity. This involves comparing amino acid changes across the spike protein of viral strains from a given month. Strains that show significantly greater changes than the average are selected as significantly altered.
These notable viral strains, which are predicted to spread more rapidly or exhibit significant immune changes, are displayed on the CoVerage platform through special graphics known as "heat maps." These visualizations allow users to quickly identify when and where major changes in the virus are occurring.
Testing the Reliability of the New Method
To evaluate the reliability of the new analysis method, researchers examined genome sequence data from known VOCs, including the omicron variant of SARS-CoV-2. The results showed that the new method could identify virus lines as VOCs up to three months before the WHO designation.
McHardy noted that variants officially recognized by the WHO showed significantly higher values in their analyses compared to less prominent variants. The numbers increased in a clear order: first for Variants under Monitoring (VUMs), then for Variants of Interest (VOIs), and finally, most strongly, for VOCs, which are considered particularly concerning.
Implications for Public Health
The findings underscore the potential of the CoVerage method to predict the emergence of health-relevant SARS-CoV-2 variants well before they reach their peak frequency or are formally identified by the WHO. This predictive capability could provide valuable time to initiate in-depth analysis required for vaccine adjustments or implement targeted measures to protect vulnerable populations.
By enhancing our ability to monitor and respond to emerging variants, tools like CoVerage play a critical role in safeguarding public health and ensuring the continued effectiveness of vaccination efforts.
Post a Comment for "Early detection of concerning SARS-CoV-2 variants through data analysis"
Post a Comment